Note
Go to the end to download the full example code.
Prony Series Viscoelastic Model (Generalized Maxwell)
import numpy as np
import matplotlib.pyplot as plt
import simcoon as sim
import os
plt.rcParams["figure.figsize"] = (18, 10)
The Prony series (generalized Maxwell) constitutive law is a rate-dependent, isotropic, linear viscoelastic model that considers thermal strains. It extends the Zener (standard linear solid) model to \(N\) Maxwell branches connected in parallel with a long-term elastic spring.
The material parameters are:
The long-term (equilibrium) Young’s modulus \(E_0\)
The long-term Poisson’s ratio \(\nu_0\)
The coefficient of thermal expansion \(\alpha\)
The number of Prony branches \(N\)
Then, for each branch \(i = 1, \ldots, N\):
The branch Young’s modulus \(E_i\)
The branch Poisson’s ratio \(\nu_i\)
The branch bulk viscosity \(\eta_{B,i}\)
The branch shear viscosity \(\eta_{S,i}\)
The constitutive law is given by:
where each viscous strain \(\boldsymbol{\varepsilon}^{v}_i\) evolves according to the Maxwell element ODE with characteristic viscosities \(\eta_{B,i}\) and \(\eta_{S,i}\).
umat_name = "PRONK" # 5 character code for the generalized Maxwell (Prony) model
E_0 = 9400.0 # Long-term Young's modulus (MPa)
nu_0 = 0.4 # Long-term Poisson's ratio
alpha = 0.0 # Coefficient of thermal expansion
n_prony = 5 # Number of Prony (Maxwell) branches
# Read branch parameters from data file
path_data = "../data"
mat_file = os.path.join(path_data, "Prony_raw.dat")
E_i, nu_i, etaB_i, etaS_i = np.loadtxt(mat_file, usecols=(0, 1, 2, 3), unpack=True)
# nstatev depends on the number of branches
nstatev = 8 + 7 * n_prony # Number of internal state variables
psi_rve = 0.0
theta_rve = 0.0
phi_rve = 0.0
solver_type = 0
corate_type = 1
# Build the properties array: [E_0, nu_0, alpha, n_prony, E_1, nu_1, etaB_1, etaS_1, ...]
props = np.array([E_0, nu_0, alpha, n_prony])
for i in range(n_prony):
props = np.append(props, [E_i[i], nu_i[i], etaB_i[i], etaS_i[i]])
path_results = "results"
pathfile = "PRONK_path.txt"
outputfile = "results_PRONK.txt"
sim.solver(
umat_name,
props,
nstatev,
psi_rve,
theta_rve,
phi_rve,
solver_type,
corate_type,
path_data,
path_results,
pathfile,
outputfile,
)
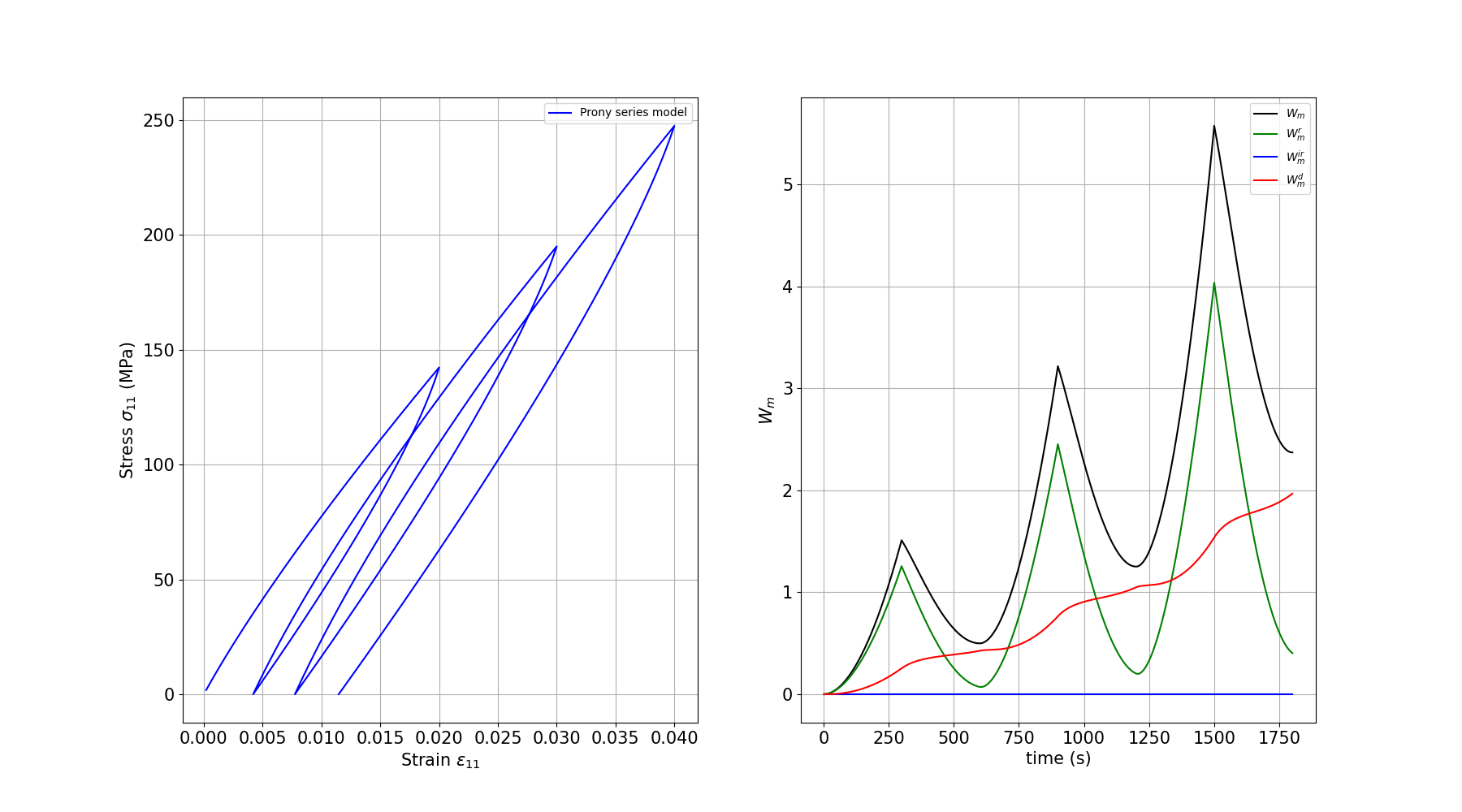
Plotting the results
We plot the stress-strain response which shows the viscoelastic behavior (rate-dependent stiffness and hysteresis from viscous dissipation).
outputfile_macro = os.path.join(path_results, "results_PRONK_global-0.txt")
fig = plt.figure()
e11, e22, e33, e12, e13, e23, s11, s22, s33, s12, s13, s23 = np.loadtxt(
outputfile_macro,
usecols=(8, 9, 10, 11, 12, 13, 14, 15, 16, 17, 18, 19),
unpack=True,
)
time, T, Q_out, r = np.loadtxt(outputfile_macro, usecols=(4, 5, 6, 7), unpack=True)
Wm, Wm_r, Wm_ir, Wm_d = np.loadtxt(
outputfile_macro, usecols=(20, 21, 22, 23), unpack=True
)
# First subplot: Stress vs Strain
ax1 = fig.add_subplot(1, 2, 1)
plt.grid(True)
plt.tick_params(axis="both", which="major", labelsize=15)
plt.xlabel(r"Strain $\varepsilon_{11}$", size=15)
plt.ylabel(r"Stress $\sigma_{11}$ (MPa)", size=15)
plt.plot(e11, s11, c="blue", label="Prony series model")
plt.legend(loc="best")
# Second subplot: Work terms vs Time
ax2 = fig.add_subplot(1, 2, 2)
plt.grid(True)
plt.tick_params(axis="both", which="major", labelsize=15)
plt.xlabel("time (s)", size=15)
plt.ylabel(r"$W_m$", size=15)
plt.plot(time, Wm, c="black", label=r"$W_m$")
plt.plot(time, Wm_r, c="green", label=r"$W_m^r$")
plt.plot(time, Wm_ir, c="blue", label=r"$W_m^{ir}$")
plt.plot(time, Wm_d, c="red", label=r"$W_m^d$")
plt.legend(loc="best")
plt.show()
Total running time of the script: (0 minutes 0.230 seconds)