Note
Go to the end to download the full example code.
Generalized Zener model (N Kelvin branches)
import numpy as np
import matplotlib.pyplot as plt
import simcoon as sim
import os
plt.rcParams["figure.figsize"] = (18, 10)
The generalized Zener model extends the standard Poynting-Thomson (Zener) model by using \(N\) Kelvin-Voigt branches in parallel with an elastic spring. This allows for a more accurate representation of the relaxation spectrum of real viscoelastic materials.
The base parameters are:
The thermoelastic Young’s modulus \(E_0\)
The thermoelastic Poisson’s ratio \(\nu_0\)
The coefficient of thermal expansion \(\alpha\)
The number of Kelvin branches \(N\)
For each branch \(i\) (\(i = 1, \ldots, N\)), four additional parameters are required:
The viscoelastic Young’s modulus \(E_i\)
The viscoelastic Poisson’s ratio \(\nu_i\)
The bulk viscosity \(\eta_{B,i}\)
The shear viscosity \(\eta_{S,i}\)
umat_name = "ZENNK" # 5 character code for the generalized Zener model
# Base material parameters
E_0 = 3000.0 # Thermoelastic Young's modulus (MPa)
nu_0 = 0.4 # Thermoelastic Poisson's ratio
alpha = 0.0 # Thermal expansion coefficient
n_kelvin = 3 # Number of Kelvin branches
# Read branch parameters from data file
mat_file = os.path.join("..", "data", "Zener_raw.dat")
E_i, nu_i, etaB_i, etaS_i = np.loadtxt(
mat_file, usecols=(0, 1, 2, 3), unpack=True
)
# nstatev depends on the number of branches
nstatev = 8 + 7 * n_kelvin
# Build the properties array: base params + branch params
props = np.array([E_0, nu_0, alpha, n_kelvin])
for i in range(n_kelvin):
props = np.append(props, [E_i[i], nu_i[i], etaB_i[i], etaS_i[i]])
psi_rve = 0.0
theta_rve = 0.0
phi_rve = 0.0
solver_type = 0
corate_type = 1
path_data = "../data"
path_results = "results"
pathfile = "ZENNK_path.txt"
outputfile = "results_ZENNK.txt"
sim.solver(
umat_name,
props,
nstatev,
psi_rve,
theta_rve,
phi_rve,
solver_type,
corate_type,
path_data,
path_results,
pathfile,
outputfile,
)
Plotting the results
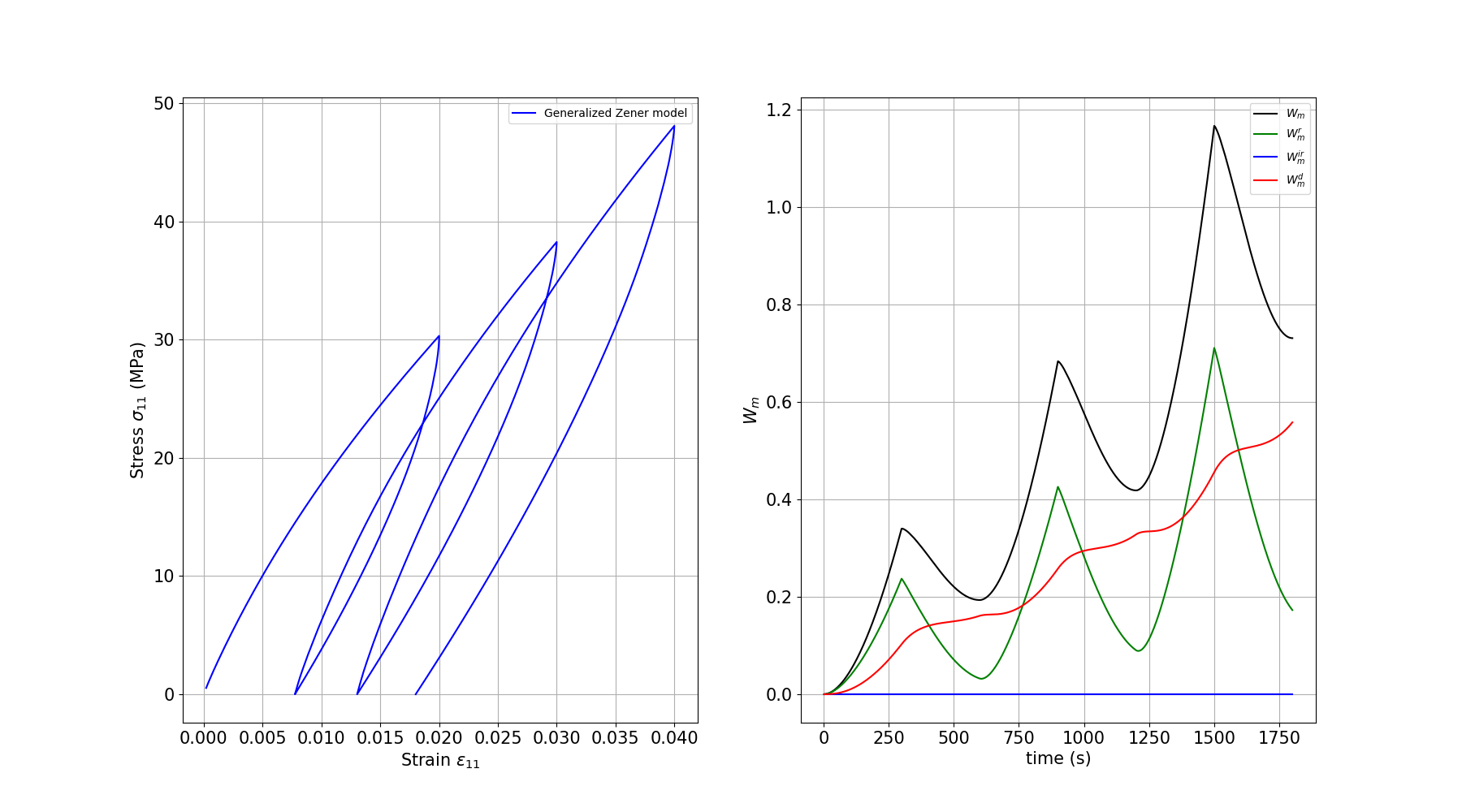
We plot the stress-strain response and the work decomposition for the generalized Zener model with multiple Kelvin branches.
outputfile_macro = os.path.join(path_results, "results_ZENNK_global-0.txt")
fig = plt.figure()
e11, e22, e33, e12, e13, e23, s11, s22, s33, s12, s13, s23 = np.loadtxt(
outputfile_macro,
usecols=(8, 9, 10, 11, 12, 13, 14, 15, 16, 17, 18, 19),
unpack=True,
)
time, T, Q_out, r = np.loadtxt(outputfile_macro, usecols=(4, 5, 6, 7), unpack=True)
Wm, Wm_r, Wm_ir, Wm_d = np.loadtxt(
outputfile_macro, usecols=(20, 21, 22, 23), unpack=True
)
# First subplot: Stress vs Strain
ax1 = fig.add_subplot(1, 2, 1)
plt.grid(True)
plt.tick_params(axis="both", which="major", labelsize=15)
plt.xlabel(r"Strain $\varepsilon_{11}$", size=15)
plt.ylabel(r"Stress $\sigma_{11}$ (MPa)", size=15)
plt.plot(e11, s11, c="blue", label="Generalized Zener model")
plt.legend(loc="best")
# Second subplot: Work terms vs Time
ax2 = fig.add_subplot(1, 2, 2)
plt.grid(True)
plt.tick_params(axis="both", which="major", labelsize=15)
plt.xlabel("time (s)", size=15)
plt.ylabel(r"$W_m$", size=15)
plt.plot(time, Wm, c="black", label=r"$W_m$")
plt.plot(time, Wm_r, c="green", label=r"$W_m^r$")
plt.plot(time, Wm_ir, c="blue", label=r"$W_m^{ir}$")
plt.plot(time, Wm_d, c="red", label=r"$W_m^d$")
plt.legend(loc="best")
plt.show()
Total running time of the script: (0 minutes 0.416 seconds)