Note
Go to the end to download the full example code.
Plasticity with isotropic hardening (thermomechanical)
import numpy as np
import matplotlib.pyplot as plt
import simcoon as sim
import os
plt.rcParams["figure.figsize"] = (18, 10)
The elastic-plastic (isotropic hardening) constitutive law implemented in simcoon is a rate independent, isotropic, von Mises type material with power-law isotropic hardening. Eight parameters are required for the thermomechanical version:
The density \(\rho\)
The specific heat \(c_p\)
The Young modulus \(E\)
The Poisson ratio \(\nu\)
The coefficient of thermal expansion \(\alpha\)
The von Mises equivalent yield stress limit \(\sigma_{Y}\)
The hardening parameter \(k\)
The hardening exponent \(m\)
The constitutive law is given by:
The updated work terms and internal heat production \(r\) are determined with the thermomechanical algorithm.
umat_name = "EPICP" # 5 character code for the elastic-plastic subroutine
nstatev = 8 # Number of internal variables
# Material parameters
rho = 4.4 # Density
c_p = 0.656 # Specific heat capacity
E = 113800.0 # Young's modulus (MPa)
nu = 0.342 # Poisson ratio
alpha = 0.86e-5 # Thermal expansion coefficient
sigma_Y = 500.0 # Yield stress (MPa)
H = 1600.0 # Hardening parameter
beta = 0.25 # Hardening exponent
psi_rve = 0.0
theta_rve = 0.0
phi_rve = 0.0
solver_type = 0
corate_type = 2
# Define the properties
props = np.array([rho, c_p, E, nu, alpha, sigma_Y, H, beta])
path_data = "../data"
path_results = "results"
# Run the simulation
pathfile = "THERM_EPICP_path.txt"
outputfile = "results_THERM_EPICP.txt"
sim.solver(
umat_name,
props,
nstatev,
psi_rve,
theta_rve,
phi_rve,
solver_type,
corate_type,
path_data,
path_results,
pathfile,
outputfile,
)
Plotting the results
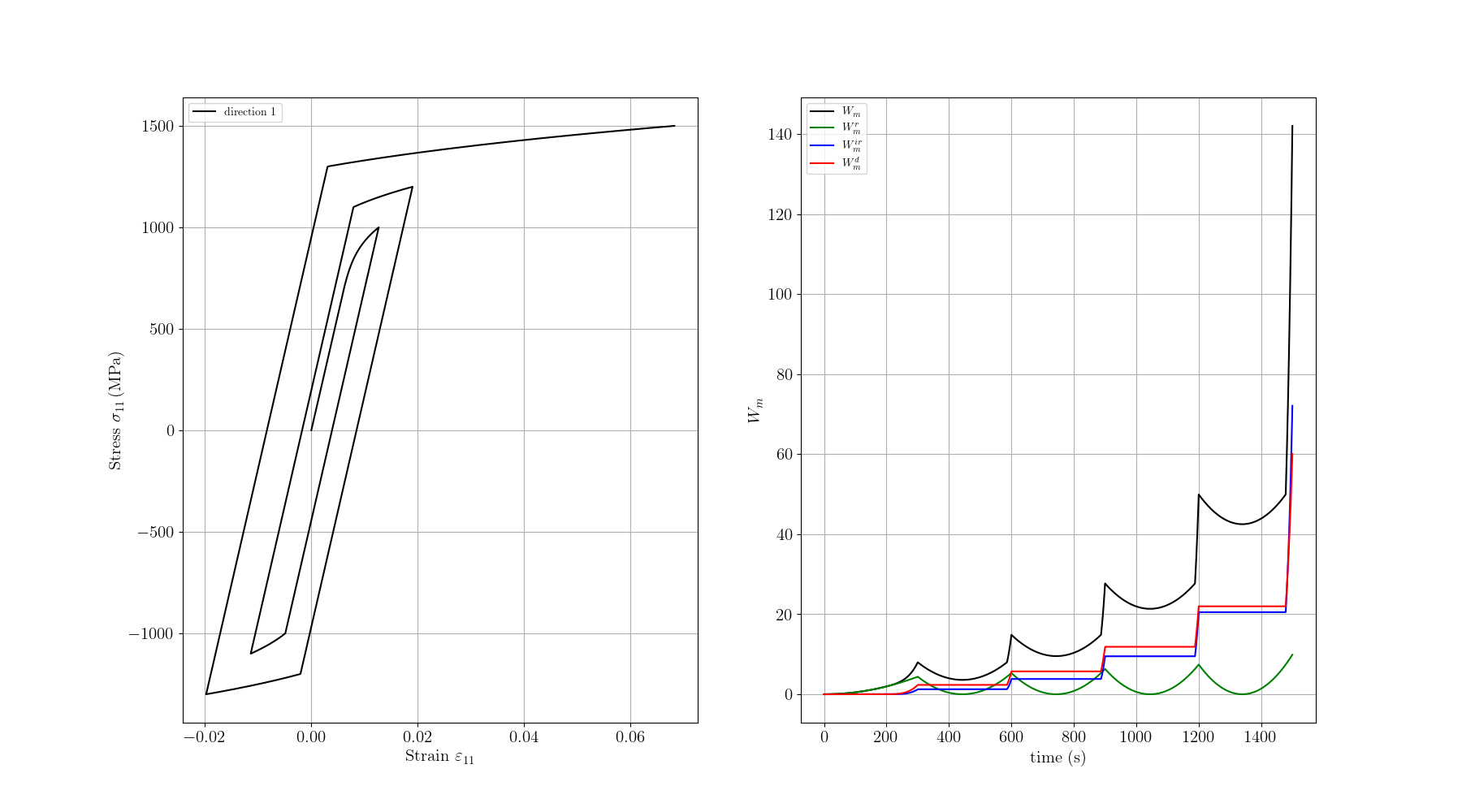
We plot the stress-strain curve, the temperature evolution, the mechanical work terms (\(W_m\), \(W_m^r\), \(W_m^{ir}\), \(W_m^d\)) and the thermal work terms (\(W_t\), \(W_t^r\), \(W_t^{ir}\)).
fig = plt.figure()
outputfile_macro = os.path.join(path_results, "results_THERM_EPICP_global-0.txt")
# Get the data
e11, e22, e33, e12, e13, e23, s11, s22, s33, s12, s13, s23 = np.loadtxt(
outputfile_macro,
usecols=(8, 9, 10, 11, 12, 13, 14, 15, 16, 17, 18, 19),
unpack=True,
)
time, T, Q, r = np.loadtxt(outputfile_macro, usecols=(4, 5, 6, 7), unpack=True)
Wm, Wm_r, Wm_ir, Wm_d, Wt, Wt_r, Wt_ir = np.loadtxt(
outputfile_macro, usecols=(20, 21, 22, 23, 24, 25, 26), unpack=True
)
# Stress vs Strain
ax = fig.add_subplot(2, 2, 1)
plt.grid(True)
plt.tick_params(axis="both", which="major", labelsize=15)
plt.xlabel(r"Strain $\varepsilon_{11}$", size=15)
plt.ylabel(r"Stress $\sigma_{11}$ (MPa)", size=15)
plt.plot(e11, s11, c="black", label="direction 1")
plt.legend(loc="best")
# Temperature vs Time
ax = fig.add_subplot(2, 2, 2)
plt.grid(True)
plt.tick_params(axis="both", which="major", labelsize=15)
plt.xlabel("time (s)", size=15)
plt.ylabel(r"Temperature $\theta$ (K)", size=15)
plt.plot(time, T, c="black", label="temperature")
plt.legend(loc="best")
# Mechanical work vs Time
ax = fig.add_subplot(2, 2, 3)
plt.grid(True)
plt.tick_params(axis="both", which="major", labelsize=15)
plt.xlabel("time (s)", size=15)
plt.ylabel(r"$W_m$", size=15)
plt.plot(time, Wm, c="black", label=r"$W_m$")
plt.plot(time, Wm_r, c="green", label=r"$W_m^r$")
plt.plot(time, Wm_ir, c="blue", label=r"$W_m^{ir}$")
plt.plot(time, Wm_d, c="red", label=r"$W_m^d$")
plt.legend(loc="best")
# Thermal work vs Time
ax = fig.add_subplot(2, 2, 4)
plt.grid(True)
plt.tick_params(axis="both", which="major", labelsize=15)
plt.xlabel("time (s)", size=15)
plt.ylabel(r"$W_t$", size=15)
plt.plot(time, Wt, c="black", label=r"$W_t$")
plt.plot(time, Wt_r, c="green", label=r"$W_t^r$")
plt.plot(time, Wt_ir, c="blue", label=r"$W_t^{ir}$")
plt.legend(loc="best")
plt.show()
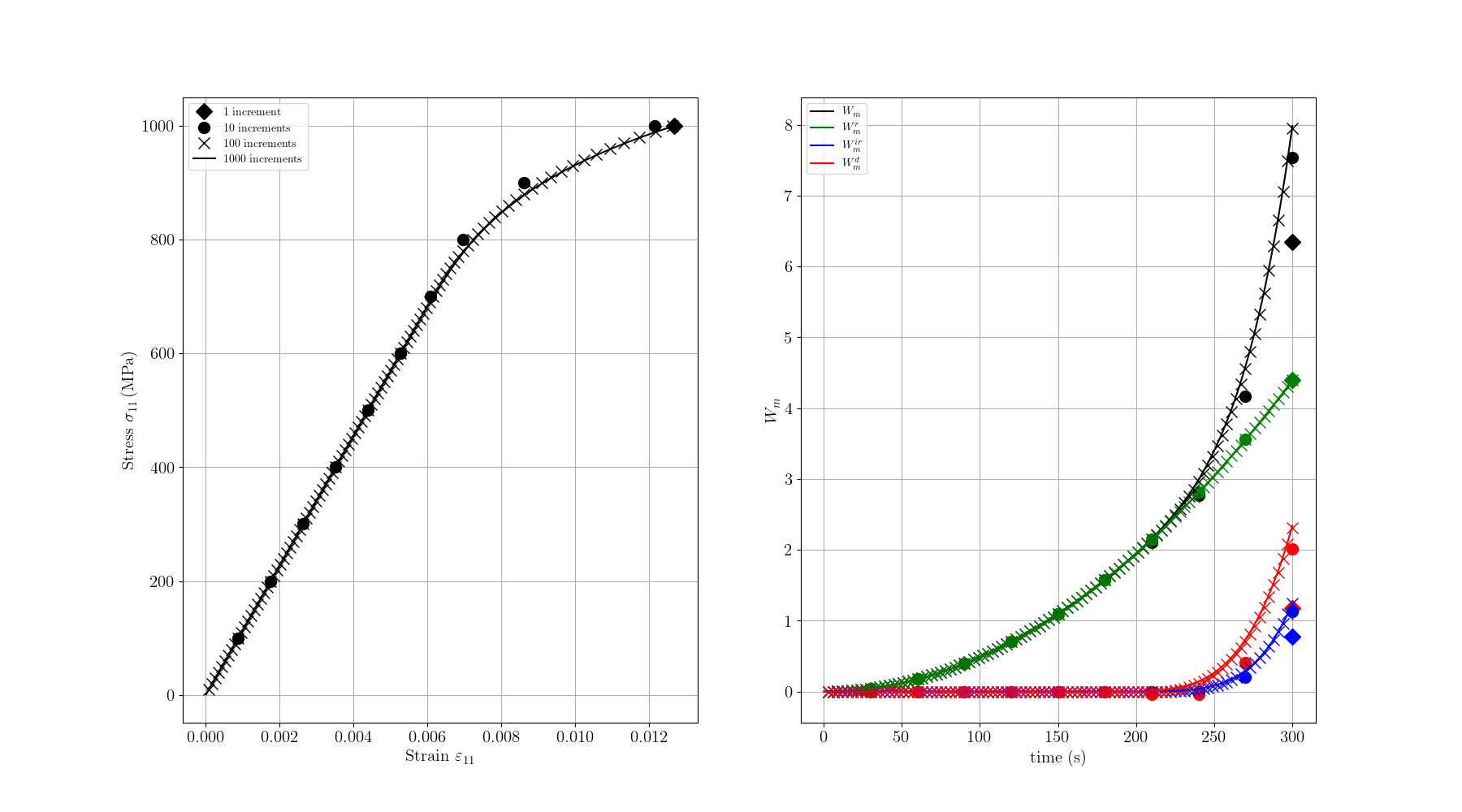
Increment size effect
Here we test the effect of the increment size on the results. We run the same simulation with 1, 10, 100, and 1000 increments.
increments = [1, 10, 100, 1000]
outputfile_globals = {}
for inc in increments:
pathfile = f"THERM_EPICP_path_{inc}.txt"
outputfile = f"results_THERM_EPICP_{inc}.txt"
sim.solver(
umat_name,
props,
nstatev,
psi_rve,
theta_rve,
phi_rve,
solver_type,
corate_type,
path_data,
path_results,
pathfile,
outputfile,
)
outputfile_globals[inc] = f"results_THERM_EPICP_{inc}_global-0.txt"
# Load data for each increment
data = []
for inc in increments:
path = os.path.join(path_results, outputfile_globals[inc])
e11, e22, e33, e12, e13, e23, s11, s22, s33, s12, s13, s23 = np.loadtxt(
path, usecols=range(8, 20), unpack=True
)
time, T, Q, r = np.loadtxt(path, usecols=range(4, 8), unpack=True)
Wm, Wm_r, Wm_ir, Wm_d, Wt, Wt_r, Wt_ir = np.loadtxt(
path, usecols=range(20, 27), unpack=True
)
data.append(
{
"e11": e11, "s11": s11, "time": time, "T": T,
"Wm": Wm, "Wm_r": Wm_r, "Wm_ir": Wm_ir, "Wm_d": Wm_d,
"Wt": Wt, "Wt_r": Wt_r, "Wt_ir": Wt_ir,
}
)
Plotting the increment size comparison
We compare the stress-strain curves, the temperature evolution, the mechanical work terms and the thermal work terms for different increment sizes.
fig = plt.figure()
markers = ["D", "o", "x", None]
labels = ["1 increment", "10 increments", "100 increments", "1000 increments"]
# Stress vs Strain
ax = fig.add_subplot(2, 2, 1)
plt.grid(True)
plt.tick_params(axis="both", which="major", labelsize=15)
plt.xlabel(r"Strain $\varepsilon_{11}$", size=15)
plt.ylabel(r"Stress $\sigma_{11}$ (MPa)", size=15)
for i, d in enumerate(data):
if markers[i] is not None:
plt.plot(d["e11"], d["s11"], linestyle="None", marker=markers[i],
color="black", markersize=10, label=labels[i])
else:
plt.plot(d["e11"], d["s11"], c="black", label=labels[i])
plt.legend(loc="best")
# Temperature vs Time
ax = fig.add_subplot(2, 2, 2)
plt.grid(True)
plt.tick_params(axis="both", which="major", labelsize=15)
plt.xlabel("time (s)", size=15)
plt.ylabel(r"Temperature $\theta$ (K)", size=15)
for i, d in enumerate(data):
if markers[i] is not None:
plt.plot(d["time"], d["T"], linestyle="None", marker=markers[i],
color="black", markersize=10, label=labels[i])
else:
plt.plot(d["time"], d["T"], c="black", label=labels[i])
plt.legend(loc="best")
# Mechanical work vs Time
ax = fig.add_subplot(2, 2, 3)
plt.grid(True)
plt.tick_params(axis="both", which="major", labelsize=15)
plt.xlabel("time (s)", size=15)
plt.ylabel(r"$W_m$", size=15)
work_colors = ["black", "green", "blue", "red"]
work_keys = ["Wm", "Wm_r", "Wm_ir", "Wm_d"]
work_labels = [r"$W_m$", r"$W_m^r$", r"$W_m^{ir}$", r"$W_m^d$"]
for i, d in enumerate(data):
for j, (wk, wc, wl) in enumerate(zip(work_keys, work_colors, work_labels)):
if markers[i] is not None:
plt.plot(d["time"], d[wk], linestyle="None", marker=markers[i],
color=wc, markersize=10)
else:
plt.plot(d["time"], d[wk], c=wc, label=wl)
plt.legend(loc="best")
# Thermal work vs Time
ax = fig.add_subplot(2, 2, 4)
plt.grid(True)
plt.tick_params(axis="both", which="major", labelsize=15)
plt.xlabel("time (s)", size=15)
plt.ylabel(r"$W_t$", size=15)
therm_keys = ["Wt", "Wt_r", "Wt_ir"]
therm_labels = [r"$W_t$", r"$W_t^r$", r"$W_t^{ir}$"]
therm_colors = ["black", "green", "blue"]
for i, d in enumerate(data):
for j, (wk, wc, wl) in enumerate(zip(therm_keys, therm_colors, therm_labels)):
if markers[i] is not None:
plt.plot(d["time"], d[wk], linestyle="None", marker=markers[i],
color=wc, markersize=10)
else:
plt.plot(d["time"], d[wk], c=wc, label=wl)
plt.legend(loc="best")
plt.show()
Total running time of the script: (0 minutes 0.612 seconds)